User Guide
1. Multiple Sequence Alignment (MSA)
To perform MSA, simply click the "Select File" button next to "Load nucleotide sequence file" in the "Consensus Sequence Generate" tab to load a viral sequence set. Viral sequence sets are stored in several different databases, including GenBank and NCBI Virus at the National Center for Biotechnology Information (NCBI, https://www.ncbi.nlm.nih.gov/ and https://www.ncbi.nlm.nih.gov/labs/virus/), the Global Initiative on Sharing All Influenza Data (GISAID, https://gisaid.org/), and Bacterial and Viral Bioinformatics Resource Center (BV-BRC, https://www.bv-brc.org/), among others. Users can retrieve viral sequence information from these platforms by downloading files in multiple formats, such as FASTA or GBK. The next step is to set up the alignment algorithm. Then, select the alignment algorithm from the menu, including AUTO (default = AUTO), FFT-NS-1, FFT-NS-2, G-INS-I, L-INS-I, and E-INS-i. Subsequently, set the consensus nucleotide threshold for consensus sequence generation in the "Bases Proportion Threshold" field (default = 0.05). Click "Run" to initiate the process. Upon completion, click "Save Alignment Seq" to save the aligned sequences and "Save Consensus Seq" to save the generated consensus sequence.
2. Single-Tube and qPCR Primer Design
Loading Files
To load an alignment, simply click the "Select File" button next to "Aligned Sequence" in the "Primer Design" tab. In addition, click the "Select File" button next to "Exclude Sequence" to input the exclusion sequence set (this field is optional), thereby enabling the primer design process to filter out potential off-target effects.
Parameter Settings
Users can select parameters in the "Options" section according to specific experimental requirements. The "Options" module provides detailed parameter settings for primer design for each sequence set, including:
1. Design oligo (default: Yes) – whether to design probes.
2. Thermodynamic formulas (default: Yes) – whether to apply thermodynamic calculations to screen candidate primers.
3. Primer pick anyway (default: No) – whether to always pick primers regardless of screening criteria.
4. Ignore gap (default: Yes) – whether to ignore Indel regions during primer design. Insertions (also called insertion mutations) involve the addition of one or more nucleotide base pairs into a DNA sequence. Deletions (also called gene deletions or deletion mutations) refer to the loss of one or more DNA segments in an organism's genome, which can occur at either the chromosomal or gene level. Collectively, insertions and deletions of bases in a genome are referred to as INDELEs.
5. Number of primers per 10,000nt (default: 10) – specifies how many primer pairs are returned.
6. Bases Proportion Threshold (default: 0.05) – sets the consensus nucleotide threshold for calculating the consensus sequence.
7. Product Size (default: 100–300 nt) – defines the desired PCR amplicon length.
8. GC% (default: 20%–80%) – specifies the GC content range of primers.
9. Tm Range (default: min 52°C, mean 56°C, max 60°C) – defines the melting temperature range of primers.
10. Primer Size (default: min 18, mean 21, max 23) – sets the length range of primers.
11. Threads (default: 4) – specifies the number of computing threads used during calculation. Primer3 supports only single-threaded computation; to enable multi-threading, VirPrimer provides a method to execute functions in parallel across multiple input values, distributing the data across processes (data parallelism). The default is set to 4, but users can customize this value according to their computational resources.
Generation of Consensus Sequences
The software generates a consensus sequence by calculating the base distribution at each position of the aligned template sequences recursively, and automatically computes the consensus according to the user-defined base proportion threshold (default p = 0.05); setting p = 0 allows the consensus sequence to represent all SNP information in the sequence set.
Execute Primer Design
Click "Run" to execute the primer design process. The software automatically uses the identified consensus sequence as input to generate primer sequences that meet the user-defined parameters.
3. Multi-Set Differential Primer Design
Loading Files
When users wish to design primers for the intersection of sequence sets (Fig. 4A), the target sequences can be merged into a single FASTA file and processed as described in the "SINGLE-TUBE AND QPCR PRIMER DESIGN" section. Conversely, to design primers for the difference between sequence sets—i.e., to align sequences from different sets, generate candidate primer sets for each set individually, and analyze the differences between these candidates—the following steps should be performed:
For each target sequence set, generate MSAs sequentially as described in the "MULTIPLE SEQUENCE ALIGNMENT" section of "Tutorial". Then, navigate to the "Primer Design" tab and click the "+" button to add the required number of sequence groups. Next, for each group, click the "Select File" button next to "Primer Design Groups" and input the MSAs for target sequence sets A, B, C, D, etc., in order. Click the "-" button to remove any unnecessary sequence groups.
Parameter Settings
Set the specific primer design parameters for each group in the "Options" section, following the procedure described in "Parameter Settings" section of "SINGLE-TUBE AND QPCR PRIMER DESIGN".
Generation of Consensus Sequences
Follow the procedure described in the "Generation of Consensus Sequences" section of "SINGLE-TUBE AND QPCR PRIMER DESIGN".
Execute Primer Design
Click "Run" to execute the primer design process. VirPrimer generates consensus sequences for each sequence group (A, B, C, D, etc.), creates candidate primer sets based on the same or different search parameters for each set, filters out primers that may cause cross-reactivity according to the respective primer design criteria, and outputs the differential primers for each group.
4. Displaying and Saving Results
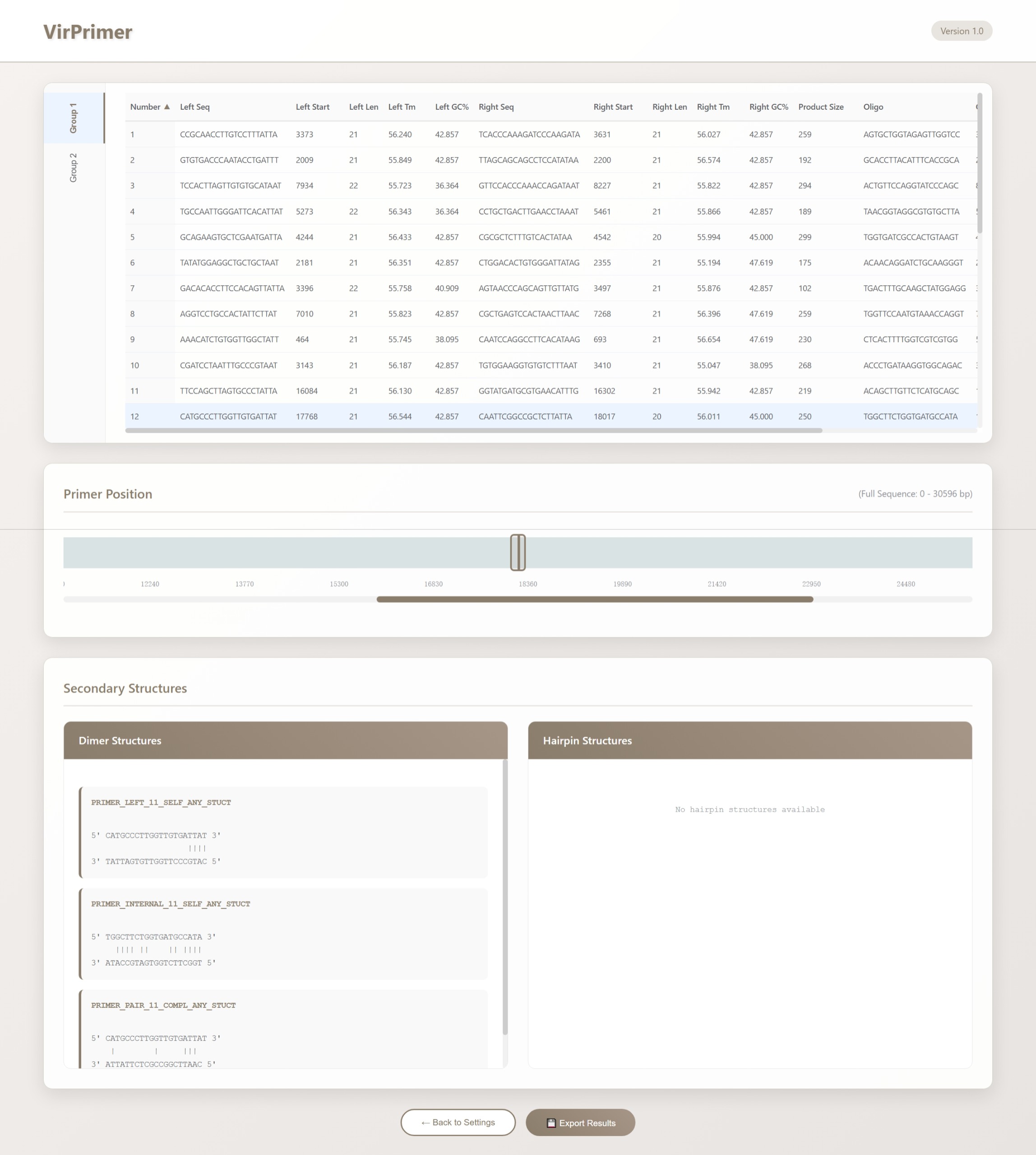
Once the computation is complete, click "Show Result" to view the output. Results are presented in tabular, graphical, and text formats. The table header provides explanations for each column, and the rows starting from the second line display the returned results.
The upper section displays the results in a table format. The columns are organized as follows:
1. Number – the identifier of the primer.
2. Left_seq – sequences of the forward primers.
3. Left_start – binding positions of the forward primers.
4. Left_len – lengths of the forward primers.
5. Left_tm – melting temperatures (Tm) of the forward primers.
6. Left_gc% – GC content percentages of the forward primers.
7. Left_any – melting temperatures of primer self-dimer formation for forward primers. (when thermodynamic calculations are applied, this column is displayed as Left_any_th)
8. Left_3' – melting temperatures of 3' self-dimer formation for forward primers. (with thermodynamic calculations, this column becomes Left_3'_th)
9. Left_hairpin – melting temperatures for hairpin formation in forward primers. (with thermodynamic calculations, this column is shown as Left_hairpin_th)
10. Right_seq – sequences of the reverse primers.
11. Right_start – binding positions of the reverse primers.
12. Right_len – lengths of the reverse primers.
13. Right_tm – Tm of the reverse primers.
14. Right_gc% – GC content percentages of the reverse primers.
15. Right_any – melting temperatures of primer self-dimer formation for reverse primers. (when thermodynamic calculations are applied, this column is displayed as Right_any_th)
16. Right_3' – melting temperatures of 3' self-dimer formation for reverse primers. (with thermodynamic calculations, this column becomes Right_3'_th)
17. Right_hairpin – melting temperatures for hairpin formation in reverse primers. (with thermodynamic calculations, this column is shown as Right_hairpin_th)
18. Oligo_seq – sequences of the probe primers.
19. Oligo_start – binding positions of the probe primers.
20. Oligo_len – lengths of the probe primers.
21. Oligo_tm – Tm of the probe primers.
22. Oligo_gc% – GC content percentages of the probe primers.
23. Oligo_any – melting temperatures of primer self-dimer formation for probe primers. (when thermodynamic calculations are applied, this column is displayed as Oligo_any_th)
24. Oligo_3' – melting temperatures of 3' self-dimer formation for probe primers. (with thermodynamic calculations, this column becomes Oligo_3'_th)
25. Oligo_hairpin – melting temperatures for hairpin formation in probe primers. (with thermodynamic calculations, this column is shown as Oligo_hairpin_th)
26. PCR_product_Size – the length of the PCR amplicon.
Click on a specific primer row to display its position within the sequence in the "Primer Position" module located at the center of the page.
Additionally, the "Dimer" module at the lower left and the "Hairpin" module at the lower right display detailed information on primer dimers and hairpin structures, respectively. If the selected primer does not form a dimer or hairpin, no corresponding result is shown. When thermodynamic calculation parameters are enabled, the modules additionally provide detailed melting parameters for dimers and hairpins, such as: Tm: 5.7°C, ΔG: -6304 cal/mol, ΔH: -43800 cal/mol, ΔS: -121 cal/mol·K.
When multiple data sets are input, VirPrimer groups and displays the results according to the sequence set, which can be switched using the group labels on the left (e.g., Group1, Group2, …). The detailed interpretation of the results is the same as described above.